
ChIP-seq
End-to-end ChIP-seq Services

- Full project: From chromatin preparation through analysis
- Library QC metrics and sequencing-ready material or FASTQ delivery
- Detailed report with QC, peak calls, and biological insights
We profile transcription factors across ChIP, CUT&Run, and CUT&Tag and guide assay selection based on target biology, antibody performance, input constraints, and failure risk. The goal is high-confidence quantitative and de-noised occupancy data with practical next-step recommendations.
3 Bulk Modalities
ChIP for TF Binding
CUT&Tag Ready
No single assay is universally best for every TF project. We evaluate target abundance, antibody behavior, sample input, and acceptable background model before selecting ChIP, CUT&Run, or CUT&Tag.
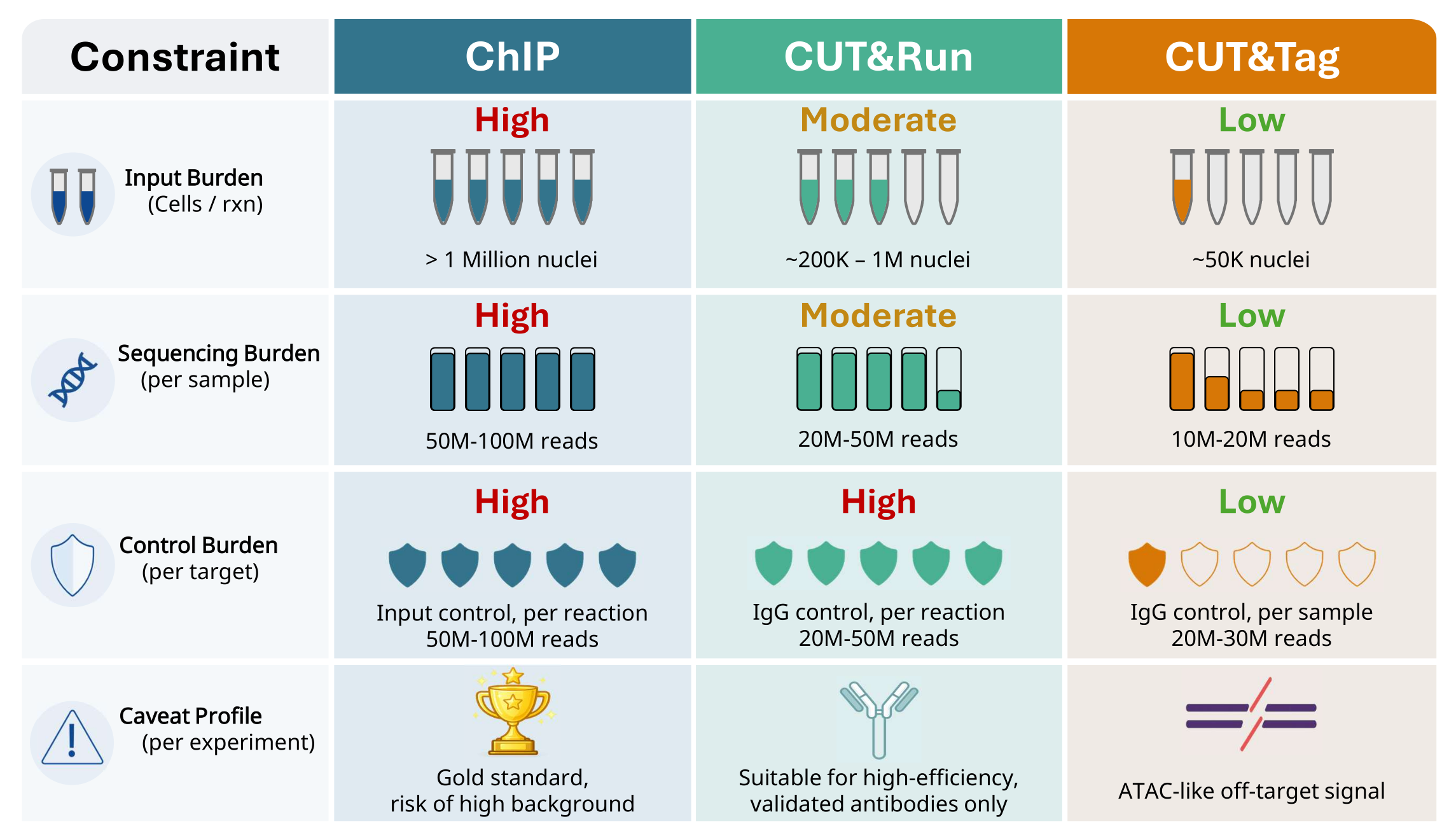
| Method | Strengths | Main Drawbacks | Typical Fit |
|---|---|---|---|
| ChIP | Gold-standard TF occupancy with conservative interpretation | Higher input burden, deeper sequencing, per-assay control requirements | Benchmark occupancy and legacy comparability |
| CUT&Tag | Lower input and sequencing burden, focused enrichment | Open-chromatin-like background can confound interpretation without controls | Input-limited or throughput-sensitive TF programs |
| CUT&Run | Lower input than ChIP in many settings | Strongly target- and antibody-dependent with potential background inflation | Bulk projects with suitable target-antibody performance |
ChIP remains the reference method for TF binding questions requiring conservative interpretation and broad comparability to legacy datasets.
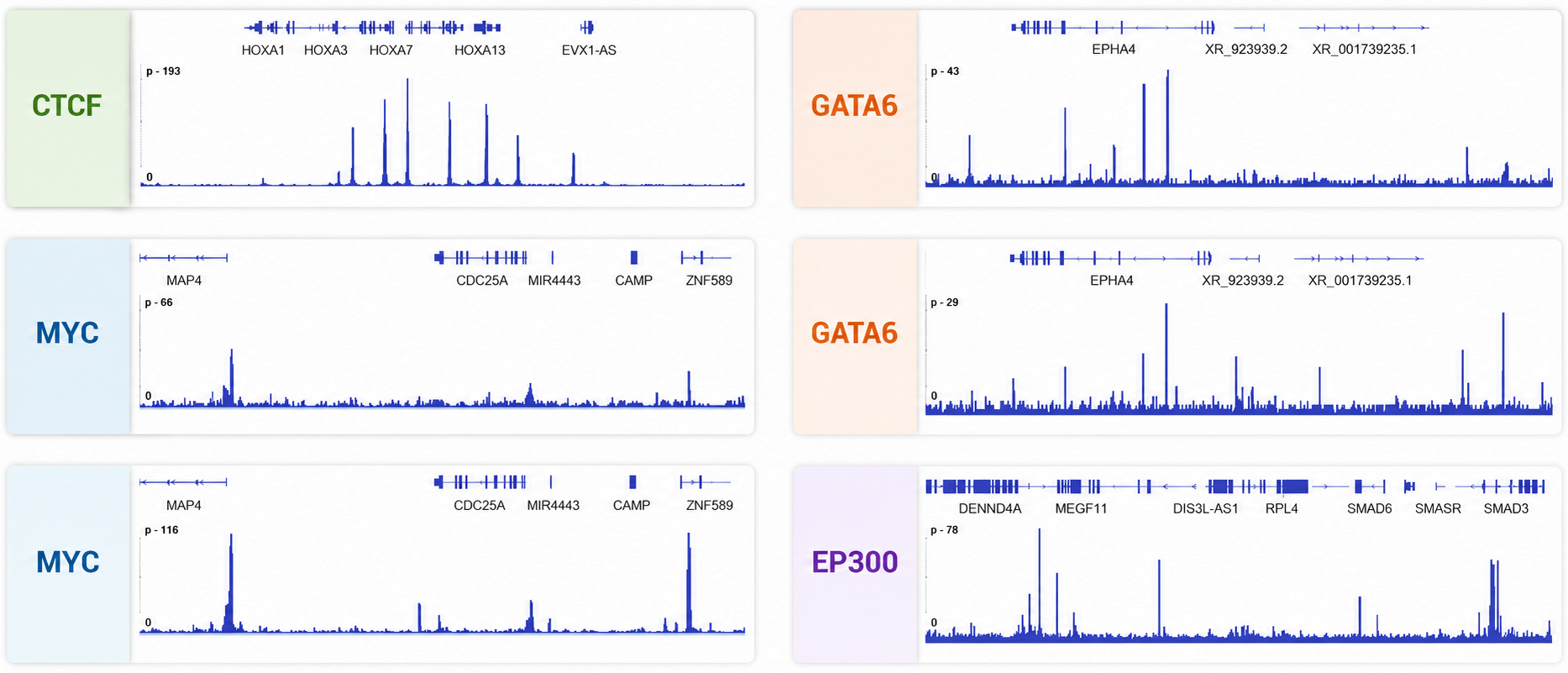
ChIP includes exogenous spike-in chromatin from a second species (typically D. melanogaster) paired with an antibody against a spike-in-species-specific target. Spike-in behavior supports both cross-condition normalization and technical QC across pulldown, library prep, and sequencing depth.
CUT&Tag commonly lowers input and sequencing requirements while producing concentrated peak signal for TF studies.
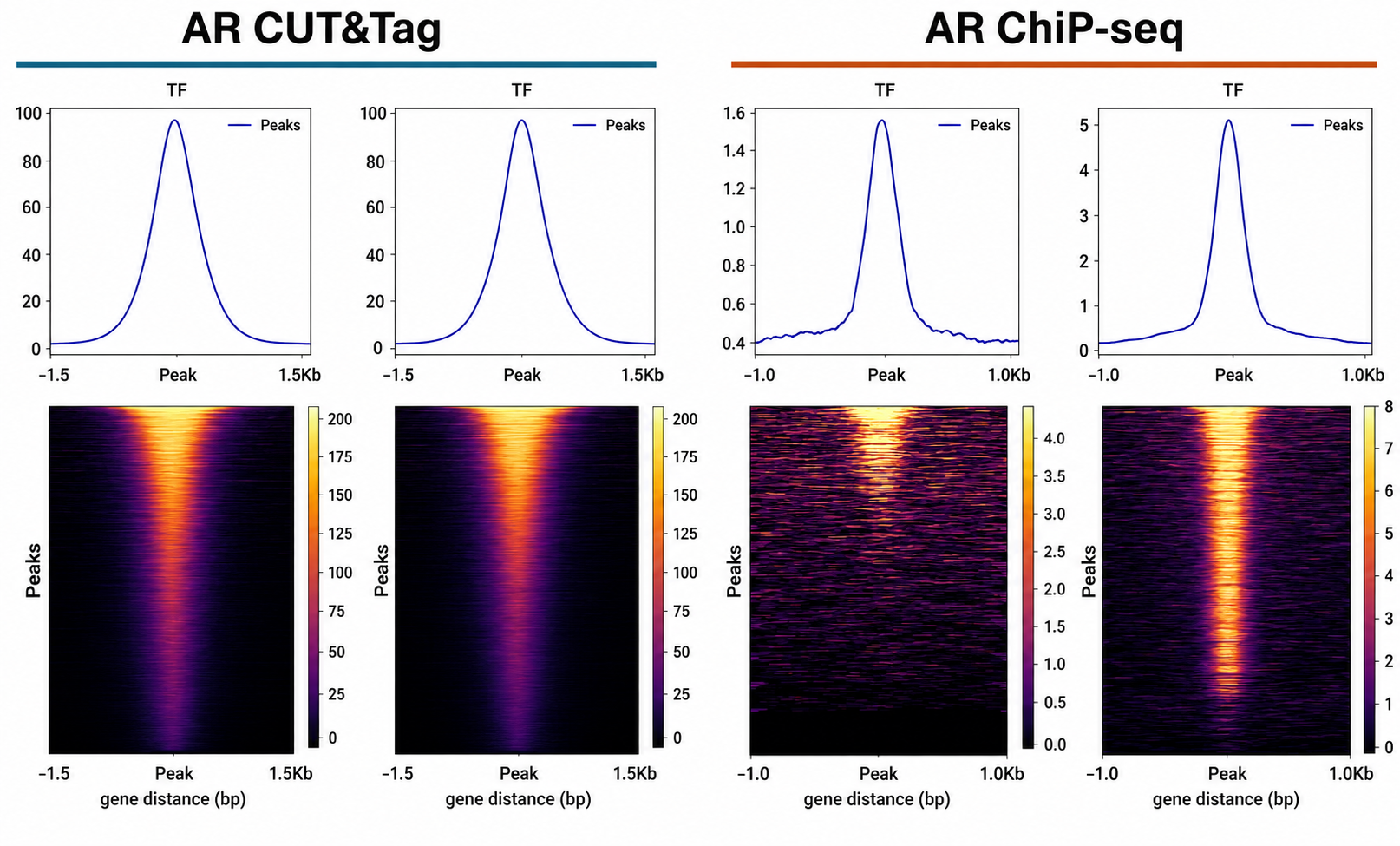
TF CUT&Tag background can resemble open chromatin. IgG/background controls, replicate concordance, and biologically anchored interpretation are standard. CUT&Tag includes spike-in nuclei rather than purified chromatin so the reference material follows the same nuclei handling and tagmentation workflow, improving both quantitative comparability and assay QC.
CUT&Run can perform strongly in suitable systems, but target abundance and antibody quality affect both digestion and pulldown outcomes.
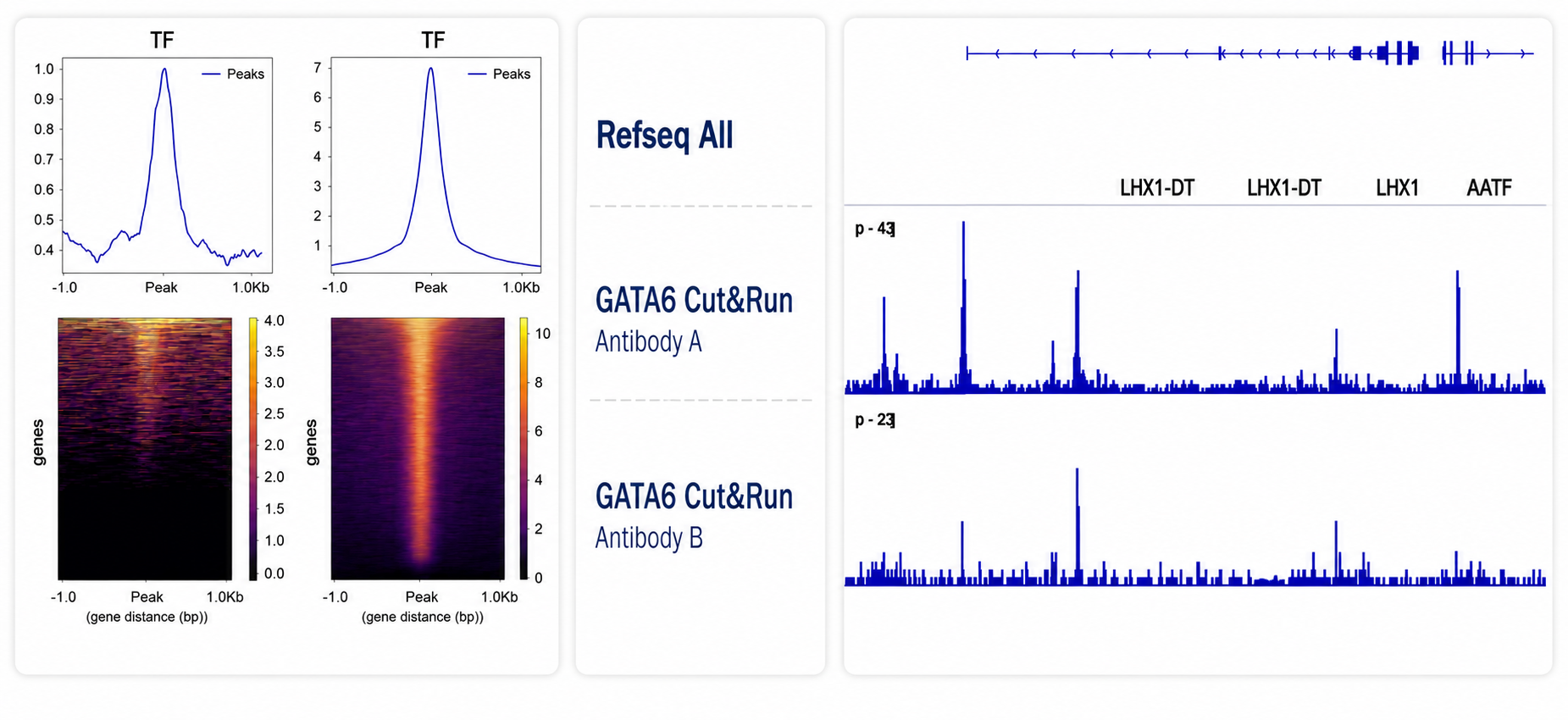
CUT&Run uses the same spike-in framework as ChIP: second-species chromatin (commonly D. melanogaster) with a species-specific antibody target. Spike-in recovery is used for normalization and as a QC indicator across digestion, pulldown, and library generation.
Single-cell TF assays are most reliable when target abundance is sufficient within each biologically relevant population. As a practical rule-of-thumb, targets near or above approximately 50,000 copies per cell are more likely to yield interpretable signal.
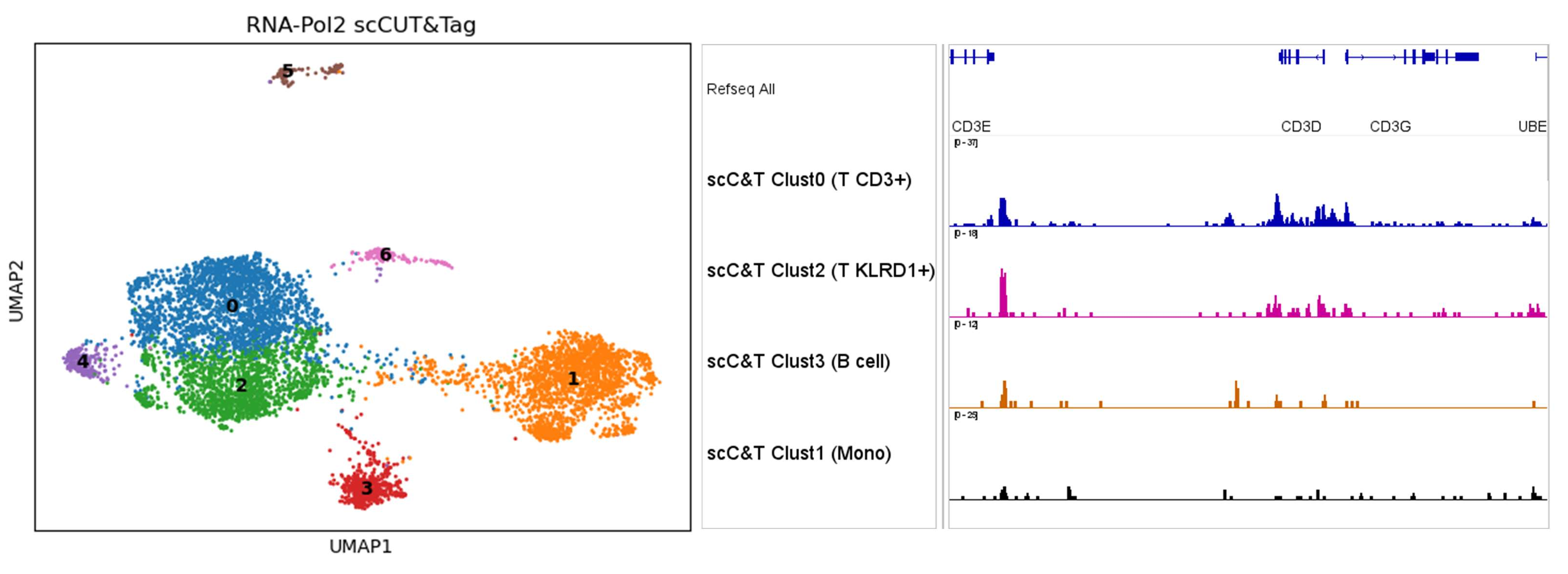
Assess target abundance, antibody constraints, and cell-population composition.
Select bulk or single-cell entry path based on interpretive risk and input constraints.
Run with assay-appropriate controls and spike-in/QC standards.
Deliver interpretation anchored to assay-specific considerations and next-step recommendations.
Start with a scientist review of your target, antibody, and sample constraints. We recommend the assay with the best balance of confidence, input feasibility, and sequencing burden for your project.